The Fraunhofer Innovation Platform for Drug Discovery and Delivery at the Hebrew University in Jerusalem, Israel, (FIP_DD@HUJI) is a joint research institution of Fraunhofer IGB and the Institute for Drug Research of the Hebrew University. The focus is on discovery and validation of new antiinfective and anti-inflammatory compounds as well as targeted nanoparticle based delivery systems for virual infections and autoimmune diseases.
The FIP_DD@HUJI is the continuation of the Fraunhofer Project Center for Drug Discovery and Delivery. It started in October 2018 and was extended in 2020 for a period of three years. The Innovation Platform was jointly headed by Prof. Dr. Steffen Rupp (Fraunhofer IGB) and Prof. Dr. Gershon Golomb (HUJI). In September 2021, Prof. Dr. Ofra Benny took over the position of director at HUJI.
Range of services
The Fraunhofer Innovation Platform combines several innovative technologies to support pharmaceutical companies in the development of new active substances at the preclinical phase. Main activities are in drug discovery and targeted delivery:
- Drug discovery/screening
- Combination of in silico-based methods and cell-based test systems for the identification of anti-infectives and immune modulators
- Identification and validation of lead structures
- Complex human-based 3D infection and tissue models with components of the immune system for validation of immune modulators and anti-infectives
- Drug transport and release
- Targeted liposomal nanocarriers for the transport of active substances into the target cells
Advantages
- Highly specific in-silico screening for agonists and antagonists of immune receptors using powerful algorithms
- Complex human-based multicellular 3D in-vitro models for validation of drug candidates and nanocarriers
- Development of nanocarriers for efficient drug transport and controlled drug release
Infrastructure: Good laboratory practice (GLP) test facility category 9: Cell-based test systems for the determination of biological parameters
Several non-clinical tests are running at our category 9 GLP unit “Cell-based test systems for the determination of biological parameters” to support R&D projects that investigate biological parameters of samples using cell-based assays.Examples are:
- Testing of bioactivity, cytotoxicity and immunogenicity of compounds using immune receptor-based assays
- Screening of TLR agonists/antagonists
- Testing of antimicrobial properties of substances or surfaces
- Detection of pyrogens and microbial residues (pathogen-associated microbial patterns, PAMPs)
- Detection of viruses
Identification of immunomodulating agents
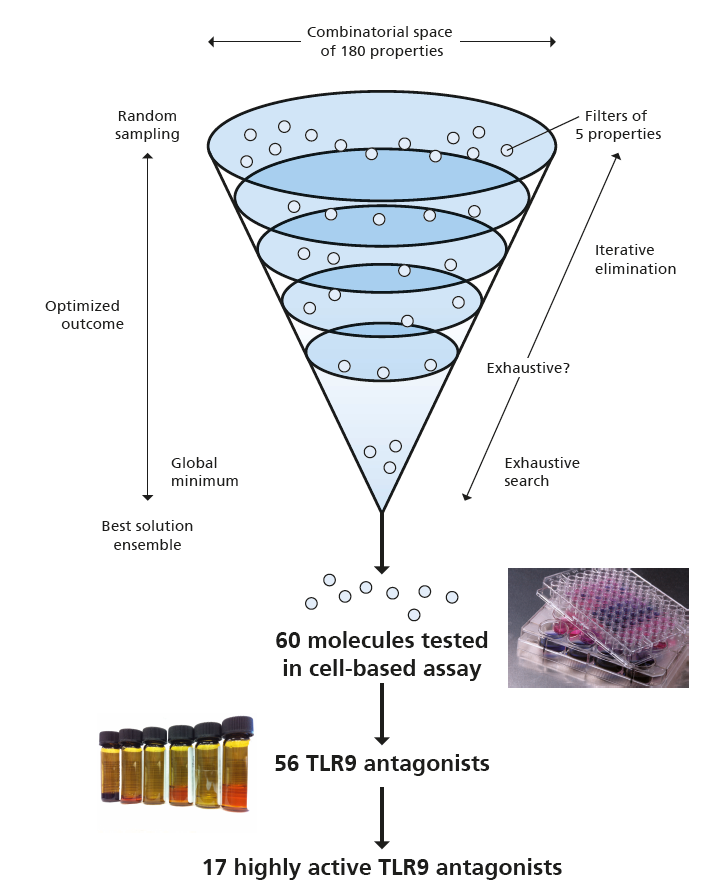
A combination of computational chemistry (on the part of HU) and a patented reporter gene assay (from IGB) is used with the aim of finding new Toll-like receptor antagonists (TLR) for modulation of the innate immune response.
During the preceding project “JRHDD – Joint Research Hub for Drug Discovery and Delivery”, 17 potential antagonists with high IC50 value were identified from 1.8 million commercially available molecules, and these were registered for patent [2].
RNA-based drugs against HSV1
Another focal point is the development of new therapeutic strategies against herpes simplex viruses (HSV). Herpes viruses cause lifelong latent infections in neuronal cells and cannot be eliminated at present. Substances such as antimicrobial peptides kill the virus but are also highly toxic to eukaryotic cells. RNA-based drugs represent a possible alternative to the antiviral drugs used to date. The use of RNA interference (RNAi) makes it possible to switch off the genes involved in the proliferation of HSV-1.
Current challenges in the application of the RNAi method arise from the fact that the inhibitory RNA is rapidly degraded by the body’s enzymes and, due to its negative charge, has extremely low membrane permeability. To overcome these problems, the Project Center uses liposomal formulations for the targeted transport of RNA molecules into the infected cell.
Nanocarriers for targeted drug transport

For encapsulation and targeted transport of active substances, e.g. siRNAs which block virus replication, liposomal formulations with optimal physico-chemical properties [3] are being developed at the Project Center. Liposomes are efficient drug delivery systems that protect the active ingredient from degradation, improve its pharmacokinetic properties and release a high concentration of active ingredient at the site of action. Targeting is performed using special navigator ligands that bind specifically to cell-type-specific receptors.
The drug-loaded liposomal transport systems (HU) are investigated and analyzed for drug release and activity using 2D and 3D cell-based test systems (IGB) [4, 5]. Various navigator peptide phospholipid formulations have shown an improved delivery of different drug candidates and their activity in 2D and 3D test systems.
ISE-CoV-2-Screen – Screening and testing therapeutic substances for the treatment of COVID-19 patients

In the anti‑corona project ISE‑CoV‑2‑Screen Fraunhofer IGB and FIP_DD@HUJI are identifying therapeutic substances for the treatment of COVID‑19 patients. The focus is on molecules that have already been approved for another indication or are at least at an advanced stage of development (repurposing of drugs).
Already approved active molecules of the pharmaceutical database DrugBank were pre-selected using computational chemistry. These have the following properties in silico:
- blocking an interaction of the docking molecules of the SARS-CoV-2 virus by binding to the so-called viral spike proteins,
- blocking an interaction at the host cell receptors by blocking the host cell ACE2 proteins, and
- blocking viral 3C-like protease, which has a key role in viral mutation and replication.
When screening 14,245 modeled compounds, 69 potential candidates were identified that possess at least two of the above multi-targeting properties. Subsequently, the pre-selected active molecules are validated in adapted in vitro test systems. These consist of human cells overexpressing the target receptor ACE2 of the virus or genetically modified reporter cells overexpressing viral spike proteins or 3C-like proteases. Initial in vitro testing confirmed inhibitory effects of the in silico drug candidates. By combining both methods, in silico design and in vitro testing, we expect that a number of promising molecules will be finally identified for further rapid development towards therapeutic medicine against SARS-CoV-2.
DRECOR – Drug delivery systems for drug candidates targeting SARS-CoV-2
In order to quickly provide therapeutics for the treatment of patients suffering from COVID‑19, drugs already approved for other indications can be screened to determine whether they are also suitable for the treatment of a SARS‑CoV‑2 infection. This approach is known as repurposing. The Fraunhofer Institute for Molecular Biology and Applied Ecology IME has already identified several drug candidates against SARS‑CoV‑2 from a repurposing library. In order to improve the efficacy of these candidates and to reduce possible side effects, the release of these drugs should take place as specifically as possible at the site of infection. Based on the criteria of biological activity, mechanism of action, pharmacokinetics, and physicochemical properties, five drug candidates were selected for further investigation of suitable formulations for application to the respiratory tract.
In parallel with the identification of the drug candidates, Fraunhofer IGB has developed tissue- and cell type-specific nanoparticular drug delivery systems for these drugs together with FIP_DD@HUJI. The partners are drawing on experience in formulating antiviral compounds against HSV-1 (herpes simplex virus) for safe transport and targeted release in human 3D tissue models. These formulations are based on liposomal release systems, which will now be investigated in combination with the drug candidates.
Literature
- Burger-Kentischer, A., Abele, I. S., Finkelmeier, D., Wiesmuller, K. H., Rupp, S. (2010) A new cell-based innate immune receptor assay for the examination of receptor activity, ligand specificity, signalling pathways and the detection of pyrogens. Journal of Immunological Methods 358: 93-103
- Zatsepin, M., Mattes, A., Rupp, S., Finkelmeier, D., Basu, A., Burger-Kentischer, A., Goldblum, A. (2016) Computational discovery and experimental confirmation of TLR9 receptor antagonist leads. J Chem Inf Model 56: 1835-1846
- Ron-Doitch, S., Sawodny, B., Kühbacher, A., David, MM., Samanta, A., Phopase, J., Burger-Kentischer, A., Griffith, M., Golomb, G., Rupp, S. (2016) Reduced cytotoxicity and enhanced bioactivity of cationic antimicrobial peptides liposomes in cell cultures and 3D epidermis model against HSV. J Control Release 229:163-71.
- Hogk, I., Kaufmann, M., Finkelmeier, D., Rupp, S., Burger-Kentischer, A. (2013) An in vitro HSV-1 reactivation model containing quiescently infected PC12 cells. BioResearch open access 2, 250-257
- Kuhbacher, A., Sohn, K., Burger-Kentischer, A., Rupp, S. (2017) Immune cell-supplemented human skin model for studying fungal infections. Methods in Molecular Biology 1508: 439-449
